XGBoost, Weighted Metrics and Hyperopt
Bayesian Tuning for Imbalanced Classification
This is a tutorial/explanation of how to set up XGBoost for imbalanced classification while tuning for imbalanced data.
There are three main sections:
- Hyperopt/Bayesian Hyperparameter Tuning
- Focal and Crossentropy losses
- XGBoost Parameter Meanings
(references are dropped as-needed)
Hyperopt
The hyperopt package is associated with Bergstra et. al.. The authors argued that the performance of a given model depends both on the fundamental quality of the algorithm as well as details of its tuning (also known as its hyper-parameters).
For the unitiated, if we have some dataset X, y and we train a model on it:
from sklearn.linear_model import ElasticNet
X = np.expand_dims(np.arange(0, 100, 1), -1)
y = 2*X + 1
lr = ElasticNet(alpha=0.5)
lr.fit(X, y)
The hyper-parameters are "meta" parameters which control the training process:
lr.get_params() # returns hyper-parameters
while parameters are (e.g. in this case) model coefficients
lr.coef_, lr.intercept_ # returns parameters
The authors of Bergstra et. al. proposed an optimization algorithm which transformed the underlying expression graph of how a performance metric is computed from hyper-parameters.
The idea (in a very summarized manner), is to take as inputs, the null prior and an experimental history of $H$ values of the loss function, and returns suggestions for which configurations to try next. Random sampling from the prior is taken as valid, and was shown to significantly increase model performance in vision-related tasks
The accompanying package which implements much of these ideas is hyperopt.
The basic steps to set this up is as follows:
# STEP 1: define a search space
SPACE = {
'param1': hp.uniform('param1', 0, 1),
'param2': hp.choice('param2', ['option1', 'option2']),
# and so on
}
# STEP 2: define an objective function
def objective(params):
# do some computation and evaluation here
loss: float # compute some loss
return {'loss': loss, 'status': STATUS_OK}
# STEP 3: evaluate the search space
best_hyperparams = fmin(
fn=objective,
max_evals=5,
space=SPACE,
algo=tpe.suggest,
trials=Trials()
)
The above needs some heavy explanation, so let's break this down:
Search Space
This is a dictionary of parameters that are used as the inputs to the optimization. Each parameter is randomly sampled (like statistic random, not "completely undefined" random) from some domain. hyperopt provides multiple methods for generating these values, but the ones I used the most are as follows:
hp.choice(label, options)
Gives a random choice from a list of options
hp.uniform(label, low, high)
Uniform float in the bounded range inclusive
The label parameter is used to to retrieve the value from the output
Objective Function
This takes in a single argument (in this case a dictionary), does some computation and returns a loss. This function must return a single dictionary with EXACTLY two entries: loss and status.
Algorithm
This is the novelty proposed in the paper. In the above I use the tree of parzen estimators (TPE), while RandomSearch and Adaptive TPE are also available
Trials
Finally, this object simply stores a list of all the parameters at a given run along with the run counter
# a dummy example showing how parameters are chosen to minimize loss
from hyperopt import STATUS_OK, Trials, fmin, hp, tpe
SPACE = {
'param1': hp.choice('param1', ['a', 'b']),
'param2': hp.uniform('param2', 0, 1)
}
# some "computation" which changes our loss
def objective(params):
loss = float('Inf')
print('params: ', params) # to show choices
if params['param1'] == 'a':
loss += 3
elif params['param1'] == 'b':
loss += 1
if params['param2'] < 0.5:
loss += 1
if params['param2'] >= 0.5:
loss += 2
return {'loss': loss, 'status': STATUS_OK}
trials = Trials()
fmin(
fn=objective,
space=SPACE,
max_evals=5,
algo=tpe.suggest,
trials=trials
)
params:
{'param1': 'a', 'param2': 0.1292582357515779}
params:
{'param1': 'b', 'param2': 0.4778423756288549}
params:
{'param1': 'b', 'param2': 0.025131981759641153}
params:
{'param1': 'a', 'param2': 0.6118336123855543}
params:
{'param1': 'a', 'param2': 0.9059446777046133}
100%|██████████| 5/5 [00:00<00:00, 269.84trial/s, best loss: inf]
{'param1': 0, 'param2': 0.1292582357515779}
Imbalanced Learning
Wang et. al proposed modification to the original XGBoost algorithm, by modification of the loss-function. Concretely, two losses were proposed as follows:
Weighted Crossentropy:
$$
Lw = -\sum{i=1}^m \left(\alpha y_i \log(\hat{y_i}) + (1-y_i)\log(1-\hat{y_i})\right)
$$
if α is greater than 1, extra loss will be counted on ’classifying 1 as 0’; On the other hand, if α is less than 1, the loss function will weight relatively more on whether data points with label 0 are correctly identified
and
Focal Loss:
$$
Lf = -\sum{i=1}^m y_i (1-\hat{y_i})^\gamma \log(\hat{y_i}) + (1-y_i)\hat{y_i}^\gamma \log(1-\hat{y}_i)
$$
If $\gamma=0$, the above becomes regular crossentropy. The paper goes into more detail on the first and second-order derivatives (since XGBoost does not implement autodif), and how it is integrated into the algorithm.
The main focus of both of the above losses is in weighting misclassification of the minority class more heavily than misclassification of the majority class.
Calling a 0 a 1 is penalized less than calling a 1 a 0 if we have substantially more 0's than 1's
The authors of the paper implemented these loss functions in imbalance-xgboost. We will not be using the entire library (since it masks too many moving parts behind high-level interfaces for my liking), but we will borrow their implementation of the Weighted Crossentropy and Focal losses:
# credit to https://github.com/jhwjhw0123/Imbalance-XGBoost
class Weight_Binary_Cross_Entropy:
'''
The class of binary cross entropy loss, allows the users to change the weight parameter
'''
def __init__(self, imbalance_alpha):
'''
:param imbalance_alpha: the imbalanced \alpha value for the minority class (label as '1')
'''
self.imbalance_alpha = imbalance_alpha
def weighted_binary_cross_entropy(self, pred, dtrain):
# assign the value of imbalanced alpha
imbalance_alpha = self.imbalance_alpha
# retrieve data from dtrain matrix
label = dtrain.get_label()
# compute the prediction with sigmoid
sigmoid_pred = 1.0 / (1.0 + np.exp(-pred))
# gradient
grad = -(imbalance_alpha ** label) * (label - sigmoid_pred)
hess = (imbalance_alpha ** label) * sigmoid_pred * (1.0 - sigmoid_pred)
return grad, hess
class Focal_Binary_Loss:
'''
The class of focal loss, allows the users to change the gamma parameter
'''
def __init__(self, gamma_indct):
'''
:param gamma_indct: The parameter to specify the gamma indicator
'''
self.gamma_indct = gamma_indct
def robust_pow(self, num_base, num_pow):
# numpy does not permit negative numbers to fractional power
# use this to perform the power algorithmic
return np.sign(num_base) * (np.abs(num_base)) ** (num_pow)
def focal_binary_object(self, pred, dtrain):
gamma_indct = self.gamma_indct
# retrieve data from dtrain matrix
label = dtrain.get_label()
# compute the prediction with sigmoid
sigmoid_pred = 1.0 / (1.0 + np.exp(-pred))
# gradient
# complex gradient with different parts
g1 = sigmoid_pred * (1 - sigmoid_pred)
g2 = label + ((-1) ** label) * sigmoid_pred
g3 = sigmoid_pred + label - 1
g4 = 1 - label - ((-1) ** label) * sigmoid_pred
g5 = label + ((-1) ** label) * sigmoid_pred
# combine the gradient
grad = gamma_indct * g3 * self.robust_pow(g2, gamma_indct) * np.log(g4 + 1e-9) + \
((-1) ** label) * self.robust_pow(g5, (gamma_indct + 1))
# combine the gradient parts to get hessian components
hess_1 = self.robust_pow(g2, gamma_indct) + \
gamma_indct * ((-1) ** label) * g3 * self.robust_pow(g2, (gamma_indct - 1))
hess_2 = ((-1) ** label) * g3 * self.robust_pow(g2, gamma_indct) / g4
# get the final 2nd order derivative
hess = ((hess_1 * np.log(g4 + 1e-9) - hess_2) * gamma_indct +
(gamma_indct + 1) * self.robust_pow(g5, gamma_indct)) * g1
return grad, hess
Application to XGBoost
Finally, we will combine the Bayesian hyperopt with the imbalanced losses and apply these to a theoretical imbalanced dataset. Before this however, we need some clarity on the xgb parameters:
booster
This determines which booster to use, can be gbtree, gblinear or dart. gbtree drops trees in order to solve over-fitting, and actually inherits from gbtree. This booster combines a large amount of regression trees with a small learning rate.
eta
This is learning rate, or how much influence the newly updated gradients affect old parameters. It reduces the influence of each individual tree and leaves space for future trees to improve the model
gamma
This is the minimum loss reduction required to further partition on a leaf node (larger gamma means more underfitting)
max_depth0 is a no-limit. In this case, this controls how deep a single decision tree is allowed to branch in any update step
subsample
This is borrowed from RandomForest, and prevents over-fitting by randomly subsampling columns to use/not-used, whilst also speeding up computations.
lambda and alpha
L2 and L1 regularization
tree_method
The ones you most commonly would use on large data are approx and hist (or if you have a GPU gpu_hist)
scale_pos_weight
This works in tandem with the imbalanced losses to upscale the penalty of misclassifying minority classes. Typically set to the number of negative instances over the number of positive instances.
Another keyword argument that we will also use is feval which allows specification of a custom evaluation metric (in our case we use a customized f1 score)
Finally, we use opt to pass in our custom objective functions
Okay let's put this all together:
# some toy data
from sklearn.datasets import make_moons
X_train, y_train = make_moons(n_samples=200, shuffle=True, noise=0.5, random_state=10)
X_test, y_test = make_moons(n_samples=200, shuffle=True, noise=0.5, random_state=10)
dtrain = xgb.DMatrix(X_train, y_train)
dtest = xgb.DMatrix(X_test, y_test)
# custom f1 evaluation metric
from sklearn.metrics import f1_score
# f1 evaluating score for XGBoost
def f1_eval(y_pred, dtrain):
y_true = dtrain.get_label()
y_pred[y_pred < 0.20] = 0
y_pred[y_pred > 0.20] = 1
err = 1-skmetrics.f1_score(y_true, np.round(y_pred))
return 'f1_err', err
import numpy as np
from sklearn.datasets import make_moons
import xgboost as xgb
from hyperopt import STATUS_OK, Trials, fmin, hp, tpe
from hyperopt.pyll import scope
# Step I: Define the search space
# here is where we use hyperopt's choice to choose between Weighted Cross Entropy and the Focal loss functoin
# as a parameter of the optimization!
wbce = Weight_Binary_Cross_Entropy(imbalance_alpha=0.5)
weighted_ce_obj = wbce.weighted_binary_cross_entropy
wf = Focal_Binary_Loss(0.5)
weighted_focal_obj = wf.focal_binary_object
SPACE = {
'n_jobs': 0,
'objective': 'binary:hinge',
'subsample': hp.uniform('subsample', 0.5, 1),
'min_child_weight': hp.uniform('min_child_weight', 1, 10),
'eta': hp.uniform('eta', 0, 1),
'max_depth': scope.int(hp.quniform('max_depth', 1, 12, 1)),
'min_split_loss': hp.uniform('min_split_loss', 0, 0.2),
'obj': hp.choice('obj', (weighted_ce_obj, weighted_focal_obj)), # hyperopt will sample one of these objectives
'num_parallel_tree': hp.choice('n_estimators', np.arange(1, 10, 1)),
'lambda': hp.uniform('lambda', 0, 1),
'alpha': hp.uniform('alpha', 0, 1),
'booster': hp.choice('booster', ['gbtree', 'gblinear', 'dart']),
'tree_method': hp.choice('tree_method', ('approx', 'hist')),
}
# Step II: Define the objective function
def objective(space):
# this is a "hack" since I want to pass obj in as
# a member of the search space
# but treat it ALONE as a keyword argument
# may increase computation time ever-so-slightly
params = {}
for k, v in space.items():
if k != 'obj' :
params[k] = v
obj = space['obj']
# train the classifier
booster = xgb.train(
params,
dtrain,
obj=obj,
feval=f1_eval # we also pass in a custom F1 evaluation metric here
)
y_pred = booster.predict(dtest)
y_pred[y_pred < 0.5] = 0
y_pred[y_pred >= 0.5] = 1
# evaluate and return
# note we want to maximize F1 and hence MINIMIZE NEGATIVE F1
return {'loss': -f1_score(y_pred, y_test), 'status': STATUS_OK}
# Step III: Optimize!
trials = Trials()
best_hyperparams = fmin(
space=SPACE,
fn=objective,
algo=tpe.suggest,
max_evals=100, # this would be 100, 500 or something higher when actually optimizing
trials=trials
)
[17:50:54] WARNING: /tmp/abs_40obctay9q/croots/recipe/xgboost-split_1659548945886/work/src/learner.cc:576:
Parameters: { "max_depth", "min_child_weight", "min_split_loss", "num_parallel_tree", "subsample", "tree_method" } might not be used.
This could be a false alarm, with some parameters getting used by language bindings but
then being mistakenly passed down to XGBoost core, or some parameter actually being used
but getting flagged wrongly here. Please open an issue if you find any such cases.
[17:50:58] WARNING: /tmp/abs_40obctay9q/croots/recipe/xgboost-split_1659548945886/work/src/learner.cc:576:
Parameters: { "max_depth", "min_child_weight", "min_split_loss", "num_parallel_tree", "subsample", "tree_method" } might not be used.
This could be a false alarm, with some parameters getting used by language bindings but
then being mistakenly passed down to XGBoost core, or some parameter actually being used
but getting flagged wrongly here. Please open an issue if you find any such cases.
100%|██████████| 100/100 [00:04<00:00, 20.51trial/s, best loss: -0.97]
# we can get the best_parameters:
print(best_hyperparams)
{'alpha': 0.1490985558271541, 'booster': 0, 'eta': 0.9253667840977927, 'lambda': 0.04797837847632137, 'max_depth': 8.0, 'min_child_weight': 1.5191535389428135, 'min_split_loss': 0.14374170690472327, 'n_estimators': 5, 'obj': 0, 'subsample': 0.986953635736163, 'tree_method': 1}
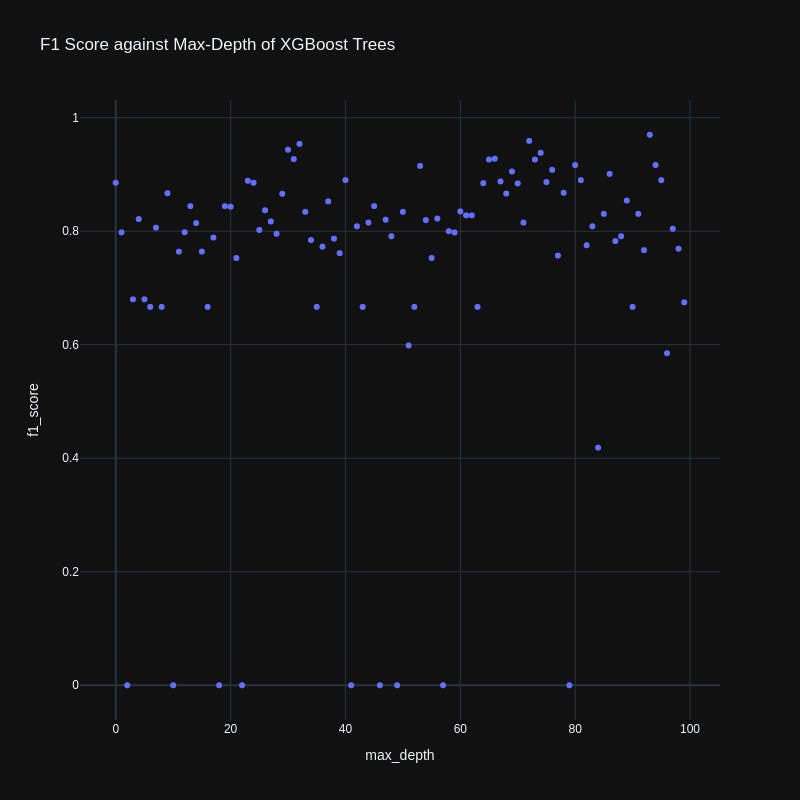
We can also plot how the F1 score varied with any of our hyperparameters:
import plotly.graph_objects as go
trials.trials[0]
fig = go.Figure(data=go.Scatter(
x=[t['misc']['idxs']['max_depth'][0] for t in trials.trials],
y=[-t['result']['loss'] for t in trials.trials],
mode='markers'
))
fig.update_layout(
xaxis=dict(title='max_depth'),
yaxis=dict(title='f1_score'),
autosize=False,
width=800,
height=800,
template='plotly_dark',
title='F1 Score against Max-Depth of XGBoost Trees'
)
fig.show()